Real-time protein quality control keeps cells healthy
March 13, 2026Scientists from the National University of Singapore (NUS) have developed a biochemical technique that captures fleeting “handshakes” between newly made proteins and the cellular helpers. These short interactions are important because they can determine whether a protein turns out healthy and useful or is faulty and in need of removal.
Cells produce vast numbers of proteins to sustain life. But building a protein is not only about assembling a chain of amino acids in the right order. As the protein chain is being produced, it must begin folding into the correct three-dimensional shape and avoid attaching to the wrong partners. When folding goes wrong, misfolded proteins can become sticky, clump together, and disrupt cellular health. Cells reduce this risk by running “quality checks” even while proteins are still being made. However, identifying the key players in this early surveillance has been challenging because their interactions with newly forming protein chains are brief and easily missed.
A method to capture short-lived interactions
To overcome this challenge, the research team led by Assistant Professor LIN Zhewang from the NUS Department of Biological Sciences developed Nascent Chain Interactor Profiling (NCIP), a method designed to capture these short-lived interactions long enough for scientists to study them. NCIP combines metabolic labelling of newly synthesised proteins with in-cell chemical crosslinking, which locks nearby molecules together. Researchers can then enrich and identify the protein partners that briefly contact nascent chains on ribosomes. This provides a systematic way to map which molecules interact with proteins during their production.
The research outcome has been published in the journal Molecular Cell.
Using NCIP, the team identified a key enzyme called TRIM25. TRIM25 can recognise misfolded proteins as they emerge from the ribosome, the cell protein making machine, and mark them for disposal before they accumulate and cause harm. It does this by attaching a small “remove me” molecular tag called ubiquitin, which directs faulty proteins to the cell’s disposal system. In other words, TRIM25 helps the cell remove problematic proteins early, before they detach from the ribosome and cause damage.
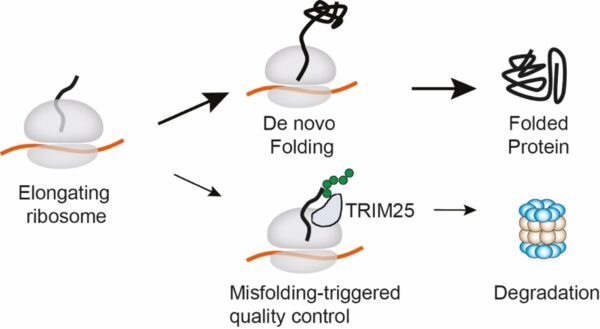
Co-translational protein quality control by TRIM25. As a new protein chain emerges from an elongating ribosome, it can either fold into its correct shape to form a functional protein, or misfold. The E3 ubiquitin ligase TRIM25 recognises misfolded nascent chains at the ribosome, tags them with ubiquitin (green dots), and directs them for degradation, helping prevent the build-up of faulty proteins in the cell. [Credit: Molecular Cell]
Relevance to diseases
This discovery matters because the build-up of misfolded and aggregated proteins is a hallmark of many neurodegenerative diseases, where abnormal proteins can progressively interfere with neuronal function. The results suggest that TRIM25 acts in a co-translational quality-control pathway that helps eliminate defective nascent proteins before they accumulate, raising the possibility that impaired TRIM25-dependent surveillance could contribute to disease susceptibility or progression.
The work may also be relevant to cancer. Rapidly proliferating tumour cells produce large quantities of proteins and often experience high “proteotoxic stress.” Enhanced protein quality control can therefore be advantageous for cancer cells survival and growth, making TRIM25 and related pathways important targets for future investigation.
Assistant Professor Lin said, “Many critical decisions about a protein’s fate happen while it is still being made. With NCIP, we can systematically capture these fleeting interactions and uncover factors, like TRIM25 that help cells detect and remove misfolded proteins early, before they become a bigger problem.”
Reference
Xiong W; Ser Z; Sobota RM; Lin Z*, “Global profiling of nascent chain interactors reveals TRIM25 as a co-translational E3 ubiquitin ligase” Molecular cell DOI: 10.1016/j.molcel.2026.02.007 Published: 2026.


